A Research Article Digest: A Conclusive Example of Evolutionarily Relevant Biological Role for G-Quadruplex Structural Motif in LINE-1 Elements

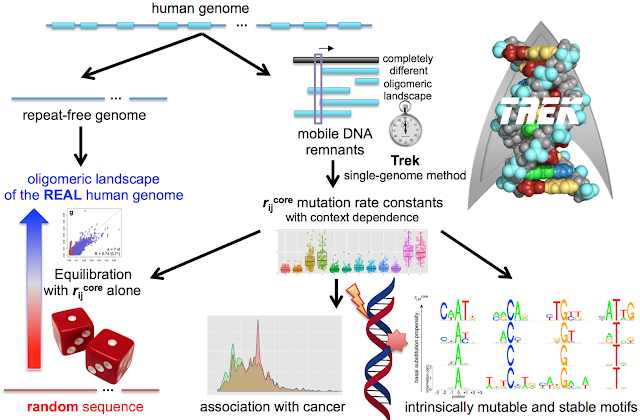
Research Article Title: G-quadruplex structures within the 3’ UTR of LINE-1 elements stimulate retrotransposition. Journal: Nature Str. Mol. Biol., 2017, aop. http://dx.doi.org/10.1038/nsmb.3367 Simple Short Explanation of the Work for Everyone: Approximately half of our genome is originated from mobile DNA elements, which are DNA segments that insert themselves at different positions in a “host” genome they infiltrate. Some mobile DNA elements increase their genomic copy numbers via a copy-and-paste mechanism, extending the length of our genome in an evolutionary timescale. In this study, we present an evidence that a specific guanine(G)-reach region in LINE-1 retrotransposons, type of mobile DNA elements that played a significant role in shaping mammalian genome architecture, has functional role in LINE-1 mobility. By a thorough dusting of the low-complexity G-rich regions in ancient LINE-1 remnants, dead at different time epochs, we revealed the or...